FragFM: Hierarchical Framework for Efficient Molecule Generation via Fragment-Level Discrete Flow Matching
Cite this work
Lee, J., Kim, S., Moon, S., Kim, H., & Kim, W. Y. (2025). FragFM: Hierarchical Framework for Efficient Molecule Generation via Fragment-Level Discrete Flow Matching. arXiv preprint arXiv:2502.15805.
@article{lee2025fragfm,
title = {FragFM: Hierarchical Framework for Efficient Molecule Generation via Fragment-Level Discrete Flow Matching},
author = {Lee, Joongwon and Kim, Seonghwan and Moon, Seokhyun and Kim, Hyunwoo and Kim, Woo Youn},
journal = {arXiv preprint arXiv:2502.15805},
year = {2025}
}
Abstract
We introduce FragFM, a novel hierarchical framework via fragment-level discrete flow matching for efficient molecular graph generation. FragFM generates molecules at the fragment level, leveraging a coarse-to-fine autoencoder to reconstruct details at the atom level. Together with a stochastic fragment bag strategy to effectively handle a large fragment space, our framework enables more efficient, scalable molecular generation. We demonstrate that our fragment-based approach achieves better property control than the atom-based method and additional flexibility through conditioning the fragment bag. We also propose a Natural Product Generation benchmark (NPGen) to evaluate the ability of modern molecular graph generative models to generate natural product-like molecules. Since natural products are biologically prevalidated and differ from typical drug-like molecules, our benchmark provides a more challenging yet meaningful evaluation relevant to drug discovery. We conduct a comparative study of FragFM against various models on diverse molecular generation benchmarks, including NPGen, demonstrating superior performance. The results highlight the potential of fragment-based generative modeling for large-scale, property-aware molecular design, paving the way for more efficient exploration of chemical space.
Contributions
Atom-based
- Chemically implausible
- Poor scaling
- Low controllability
Fragment-based
- Limited fragment library
- Poor scaling
- Hard to recover whole graph
FragFM
oursThe first fragment-level graph flow matching that explores meaningful chemical space at scale.
Method
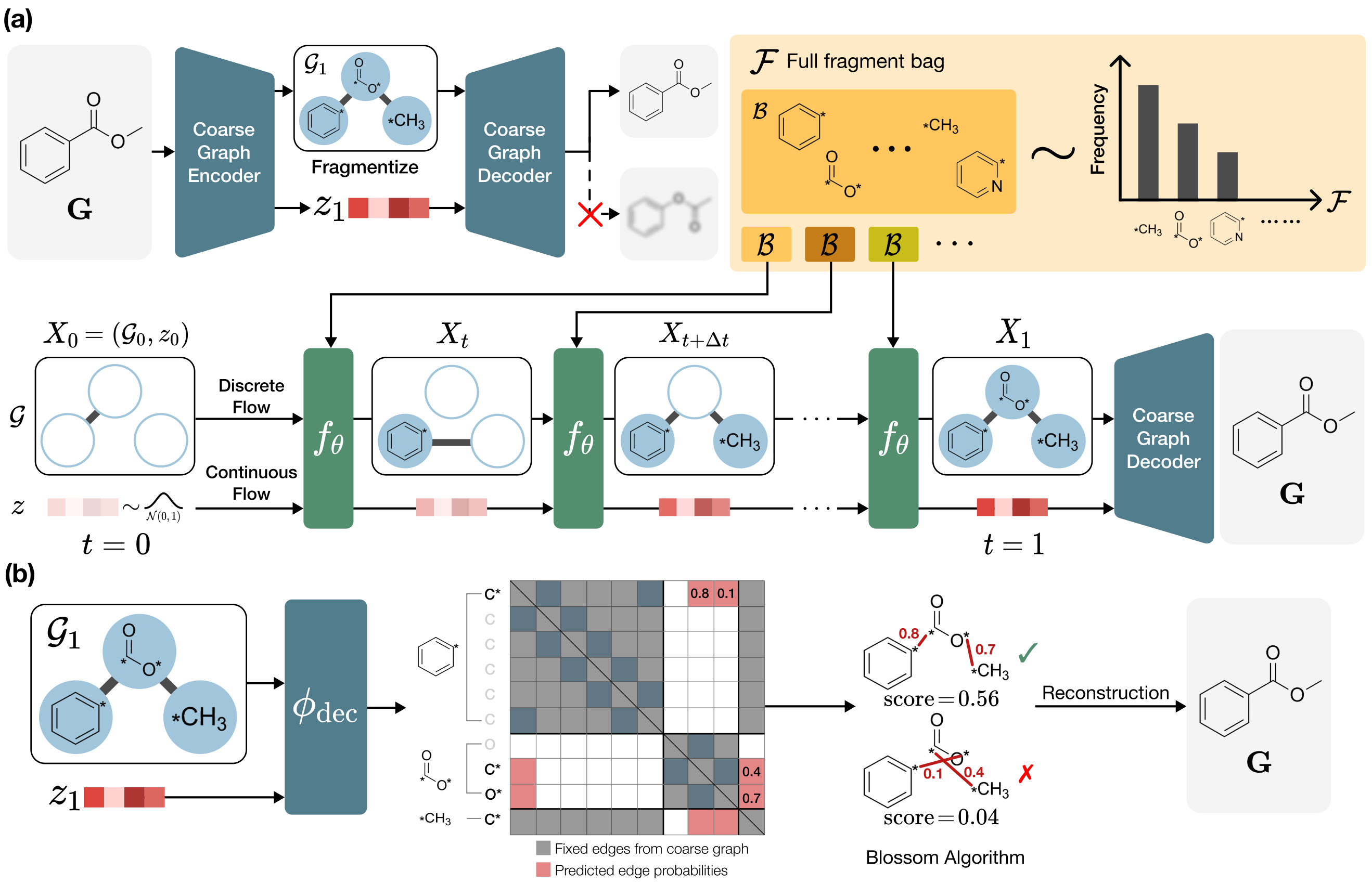
FragFM rests on two ideas. (i) A coarse-to-fine autoencoder compresses an atom-level graph G into a much smaller fragment-level graph 𝒢 plus a single continuous latent z that carries the atom-level connectivity. (ii) On that fragment-level graph, discrete flow matching (DFM) generates fragment types; to make the huge fragment vocabulary tractable, training and sampling are restricted to a stochastic fragment bag 𝓑 ⊂ 𝓕, optimized with an Info-NCE objective.
Coarse-to-Fine Autoencoder
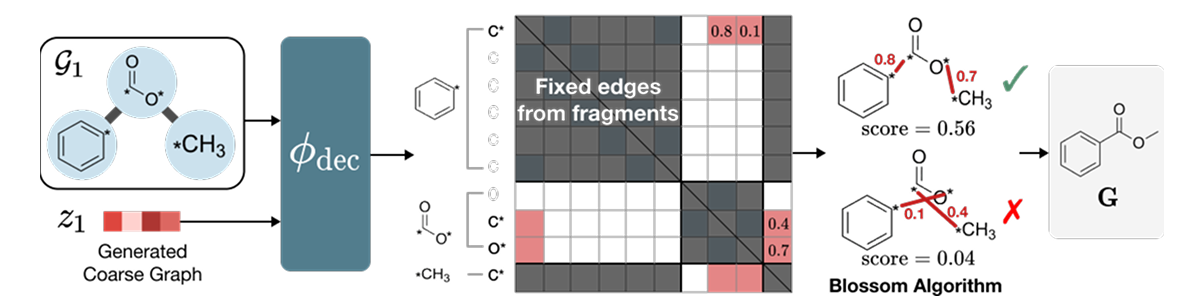
The encoder first applies a deterministic fragmentation rule (BRICS) to split G into fragments, then a graph neural network φenc encodes both into a latent z that remembers which atoms were bonded across fragment cuts. The decoder φdec scores candidate inter-fragment atom pairs and feeds those scores into the Blossom matching algorithm to recover exact atom-level edges E.
| Dataset | Bond | Graph |
|---|---|---|
| MOSES | 99.99 | 99.93 |
| GuacaMol | 99.98 | 99.42 |
| ZINC250k | 99.64 | 98.71 |
| NPGen | 99.71 | 97.43 |
Fragment-Level Flow Matching with Stochastic Bag
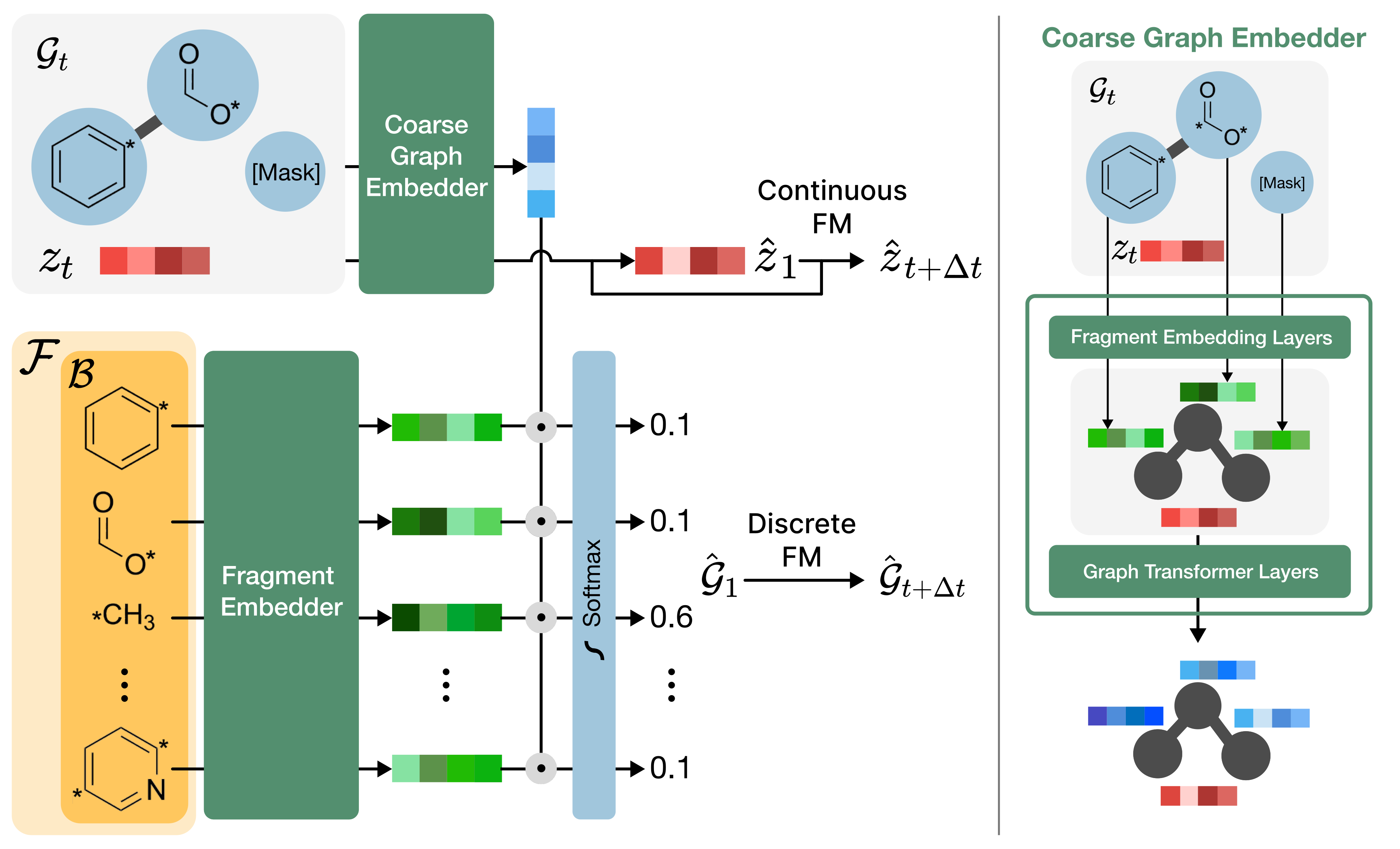
On the coarse graph X = (𝒢, z), FragFM runs discrete flow matching (DFM) over fragment types and continuous flow matching over the latent. To avoid a softmax over the full fragment vocabulary 𝓕 (which can exceed 10⁵ fragments), each step samples a bag 𝓑 ⊂ 𝓕 of size N and restricts the denoiser's softmax to 𝓑. A neural density-ratio estimator fθ(Xt, x) is trained with an Info-NCE loss against one positive + N−1 negative fragments:
At inference, the trained fθ defines an in-bag posterior over 𝓑, which the sampler integrates into a one-step Euler transition kernel:
Benchmarks
NPGen — A Natural Product Benchmark
658,566 natural products from COCONUT, average 35.0 heavy atoms (vs. 21.7 MOSES, 27.9 GuacaMol). Evaluation adds NP-likeness and NP-Classifier divergences that reflect biological functionality rather than saturated distributional overlap.
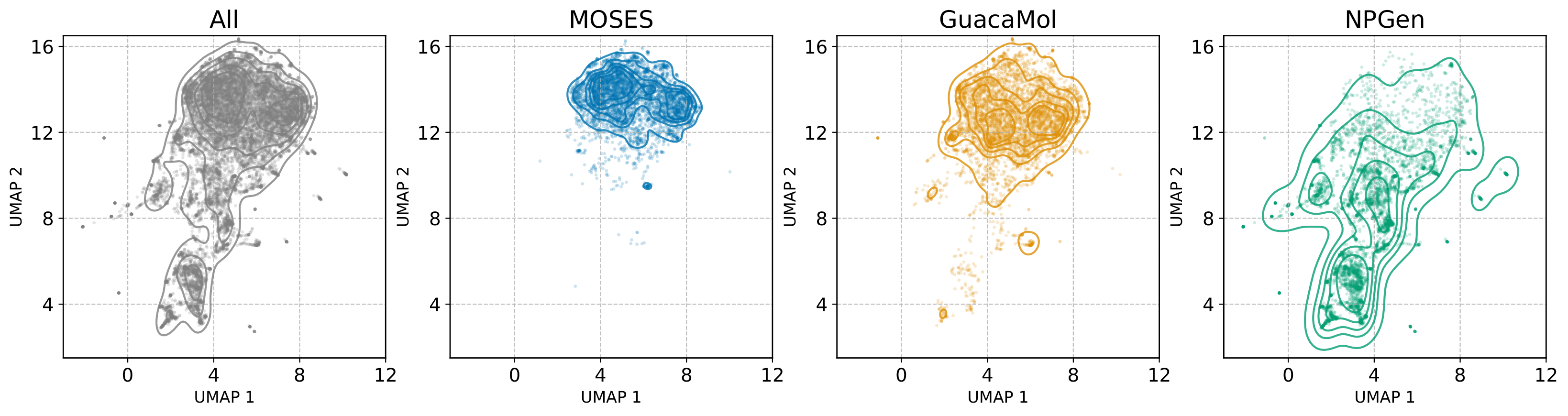
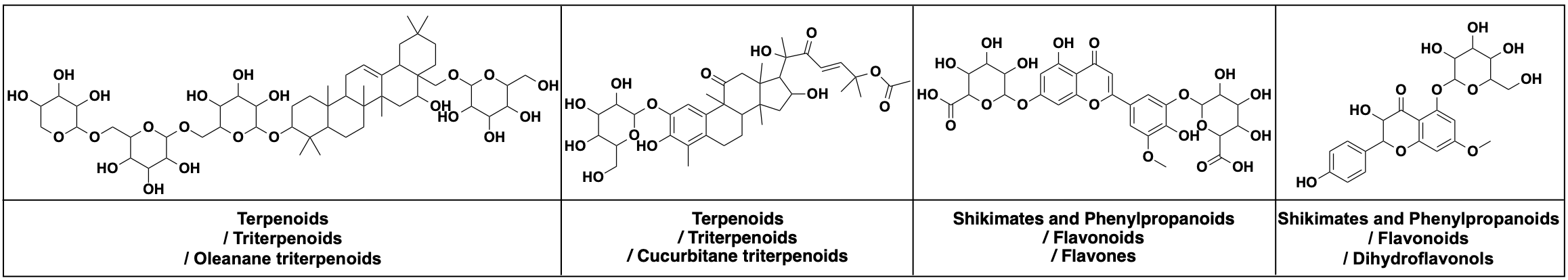
Quantitative Results
FragFM is a graph generative model, evaluated against atom- and fragment-level graph baselines on four standard molecular-generation benchmarks — matching or setting SOTA across all of them. autoregressive / sequence and one-shot diffusion / flow baselines are shown for context. Bold = best; underline = second-best.
| Model | Valid ↑ | Unique ↑ | Novel ↑ | Filters ↑ | FCD ↓ | SNN ↑ |
|---|---|---|---|---|---|---|
| Training set | 100.0 | 100.0 | – | 100.0 | 0.48 | 0.59 |
| GraphINVENT | 96.4 | 99.8 | – | 95.0 | 1.22 | 0.54 |
| JT-VAE | 100.0 | 100.0 | 99.9 | 97.8 | 1.00 | 0.53 |
| DiGress | 85.7 | 100.0 | 95.0 | 97.1 | 1.19 | 0.52 |
| DisCo | 88.3 | 100.0 | 97.7 | 95.6 | 1.44 | 0.50 |
| Cometh | 90.5 | 100.0 | 96.4 | 97.2 | 1.44 | 0.51 |
| Cometh-PC | 90.5 | 99.9 | 92.6 | 99.1 | 1.27 | 0.54 |
| DeFoG | 92.8 | 99.9 | 92.1 | 98.9 | 1.95 | 0.55 |
| FragFM (ours) | 99.8 | 100.0 | 87.1 | 99.1 | 0.58 | 0.56 |
| Model | Val. ↑ | V.U. ↑ | V.U.N. ↑ | KL Div. ↑ | FCD ↑ |
|---|---|---|---|---|---|
| Training set | 100.0 | 100.0 | – | 99.9 | 92.8 |
| MCTS | 100.0 | 100.0 | 95.4 | 82.2 | 1.5 |
| DiGress | 85.2 | 85.2 | 85.1 | 92.9 | 68.0 |
| DisCo | 86.6 | 86.6 | 86.5 | 92.6 | 59.7 |
| Cometh | 94.4 | 94.4 | 93.5 | 94.1 | 67.4 |
| Cometh-PC | 98.9 | 98.9 | 97.6 | 96.7 | 72.7 |
| DeFoG | 99.0 | 99.0 | 97.9 | 97.7 | 73.8 |
| FragFM (ours) | 99.7 | 99.3 | 95.0 | 97.4 | 85.8 |
| Model | Valid ↑ | NSPDK ↓ | FCD ↓ |
|---|---|---|---|
| Training set | – | 0.0001 | 0.062 |
| GraphAF | 67.92 | 0.0432 | 16.128 |
| GraphDF | 89.72 | 0.1737 | 33.899 |
| MolHF | 94.75 | 0.0709 | 22.230 |
| GDSS | 97.12 | 0.0192 | 14.032 |
| GSDM | 92.57 | 0.0168 | 12.435 |
| GruM | 98.32 | 0.0023 | 2.235 |
| SwinGNN | 86.16 | 0.0047 | 4.398 |
| DiGress | 94.98 | 0.0021 | 3.482 |
| GGFlow | 99.63 | 0.0010 | 1.455 |
| FragFM (ours) | 99.81 | 0.0002 | 0.630 |
| Model | Val. ↑ | Unique ↑ | Novel ↑ | NP Score KL Div. ↓ |
NP Class KL Div. ↓ | FCD ↓ | ||
|---|---|---|---|---|---|---|---|---|
| Pathway | Superclass | Class | ||||||
| Training set | 100.0 | 100.0 | – | 0.0006 | 0.0002 | 0.0028 | 0.0094 | 0.01 |
| GraphAF | 79.1 | 63.6 | 95.6 | 0.8546 | 0.9713 | 3.3907 | 6.6905 | 25.11 |
| JT-VAE | 100.0 | 97.2 | 99.5 | 0.5437 | 0.1055 | 1.2895 | 2.5645 | 4.07 |
| HierVAE | 100.0 | 81.5 | 97.7 | 0.3021 | 0.4230 | 0.5771 | 1.4073 | 8.95 |
| DiGress | 85.4 | 99.7 | 99.9 | 0.1957 | 0.0229 | 0.3370 | 1.0309 | 2.05 |
| DeFoG | 85.9 | 98.4 | 99.2 | 0.1550 | 0.1252 | 0.4134 | 1.3597 | 4.46 |
| FragFM (ours) | 98.0 | 99.0 | 95.4 | 0.0374 | 0.0196 | 0.1482 | 0.3570 | 1.34 |
NPGen Samples Across Models
Click a model to browse its NPGen samples — across small (≤30 heavy atoms) and large (31–60) molecules.
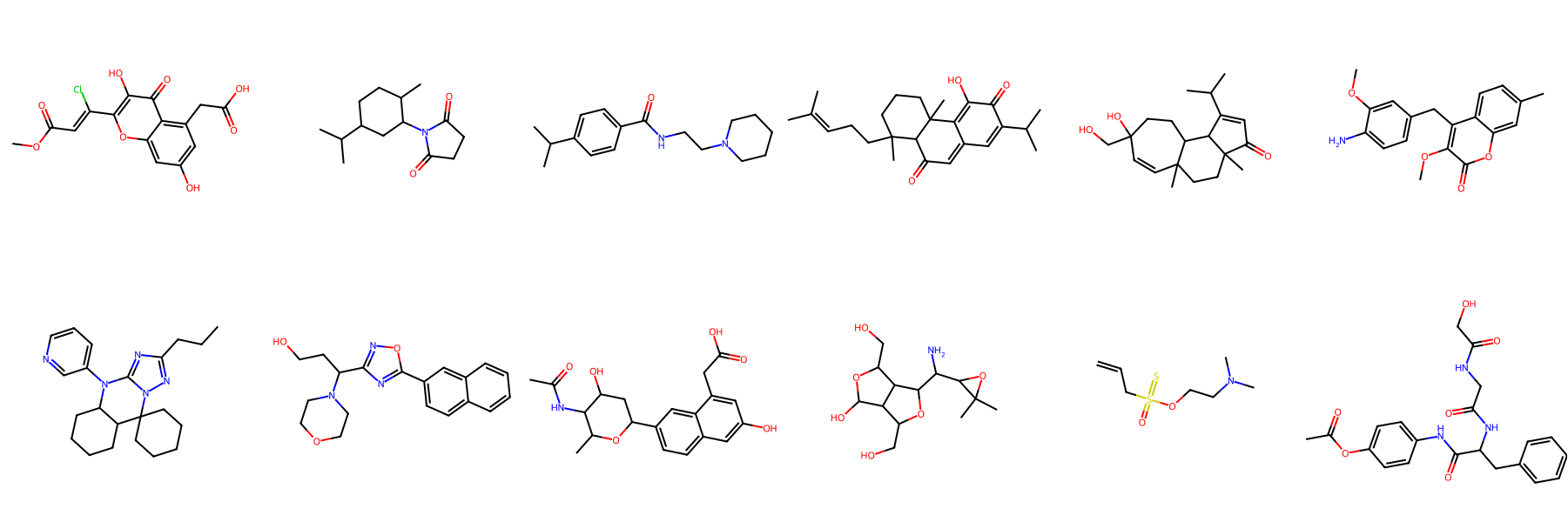
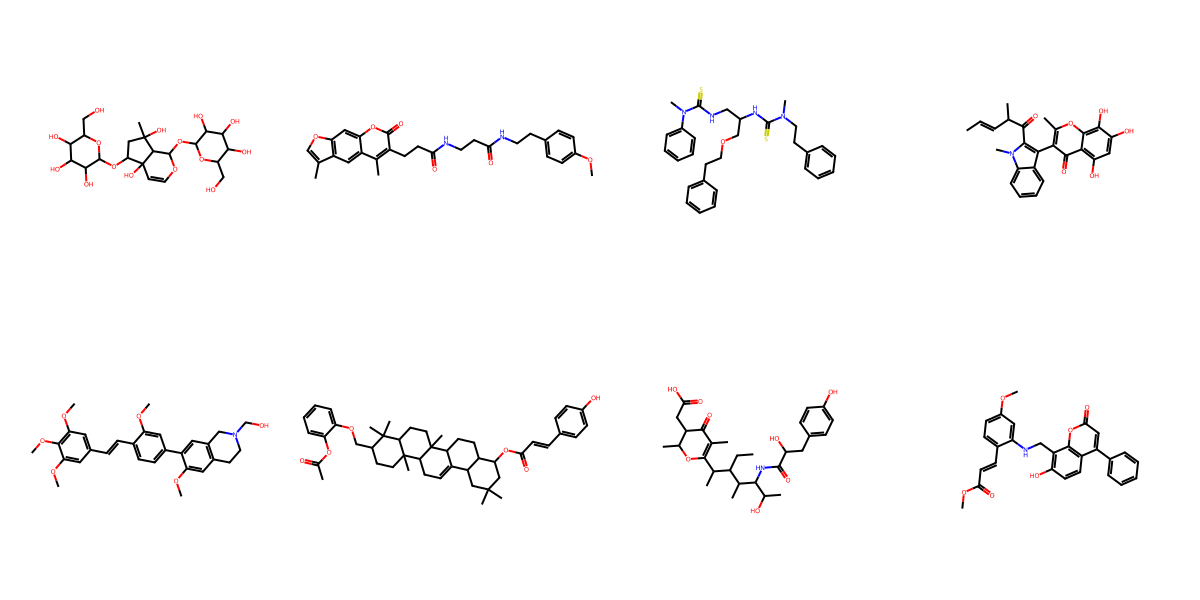
Generated Sample Gallery
Random valid molecules generated by FragFM on the standard drug-like benchmarks MOSES and GuacaMol.
Conditional Generation
Property-Controlled Generation
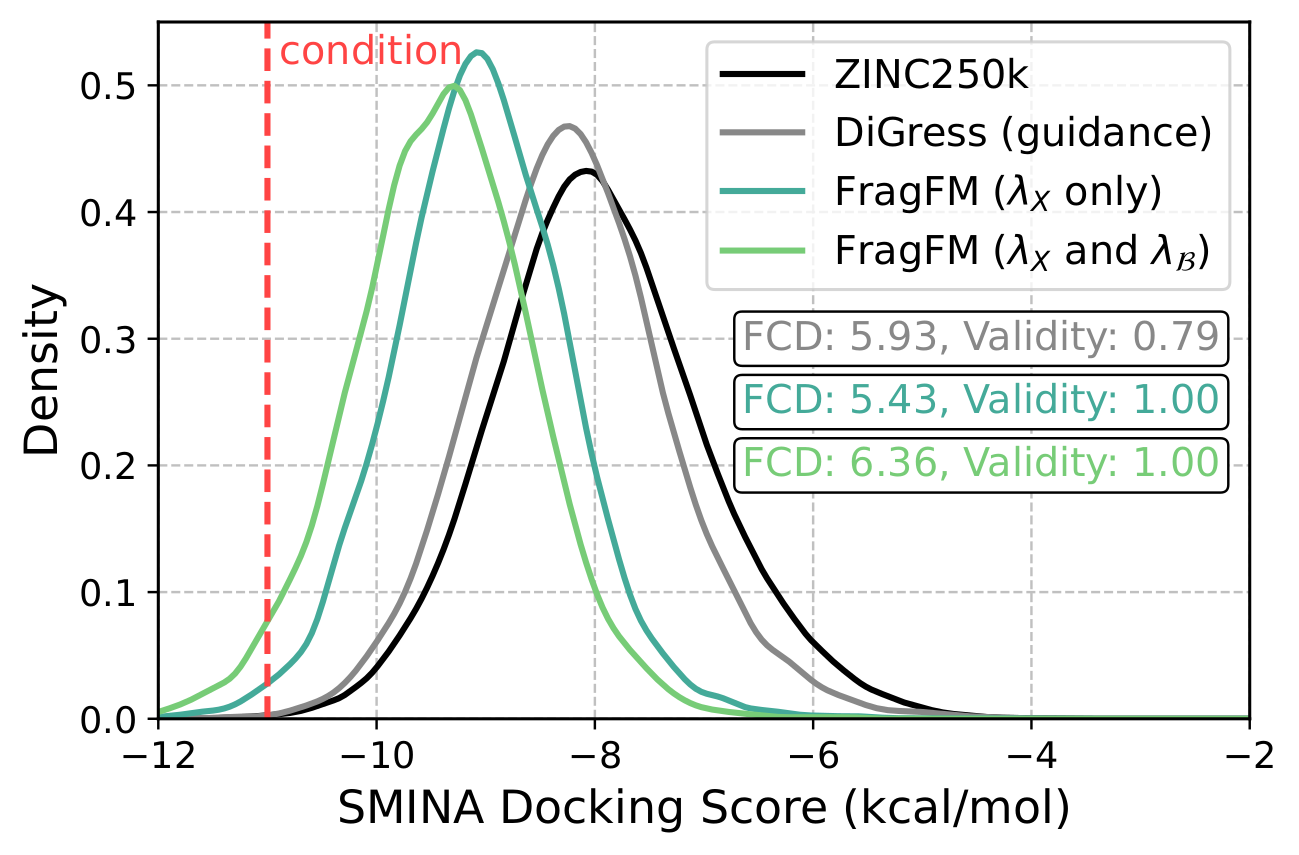
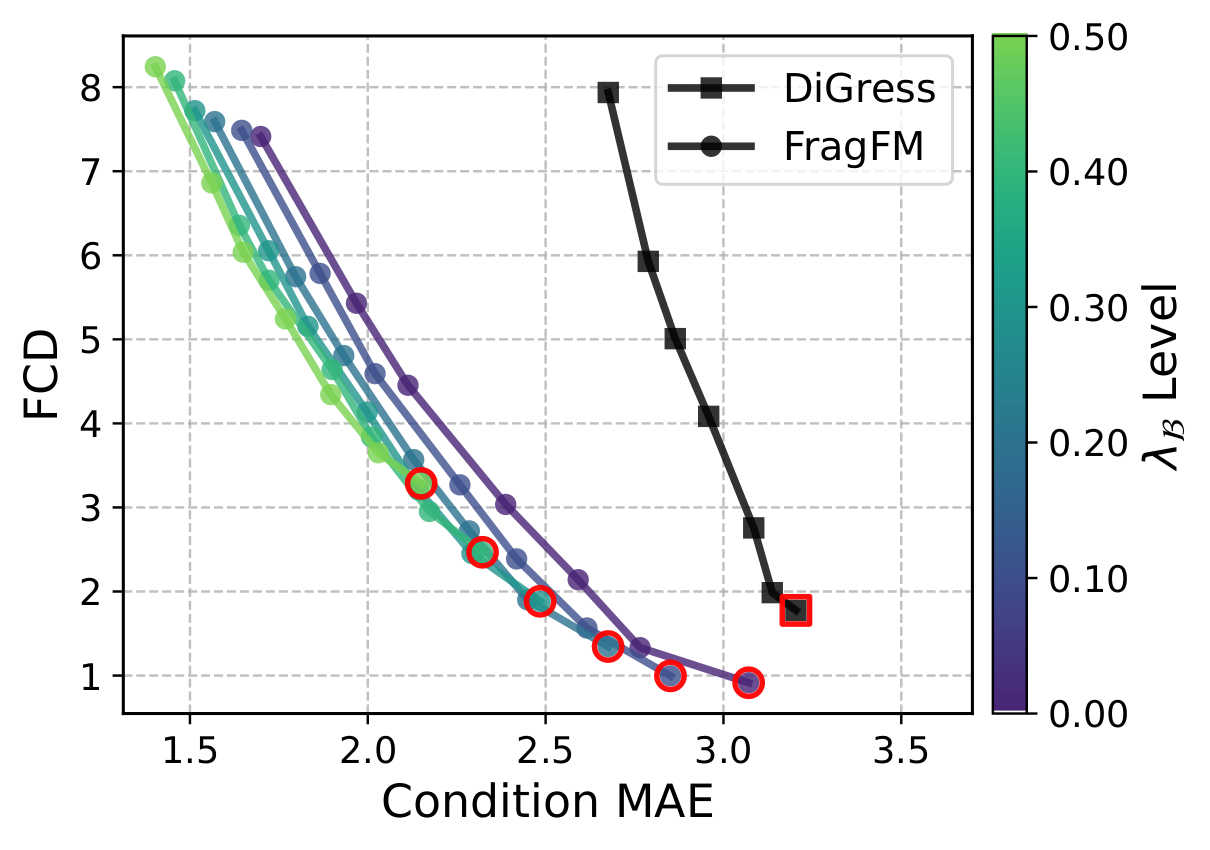
FragFM exposes two guidance knobs: standard classifier guidance λX that pulls the flow trajectory toward a target property c, and fragment-bag reweighting λ𝓑 that biases the fragment bag 𝓑 itself — up-weighting fragments whose predicted property matches the target.
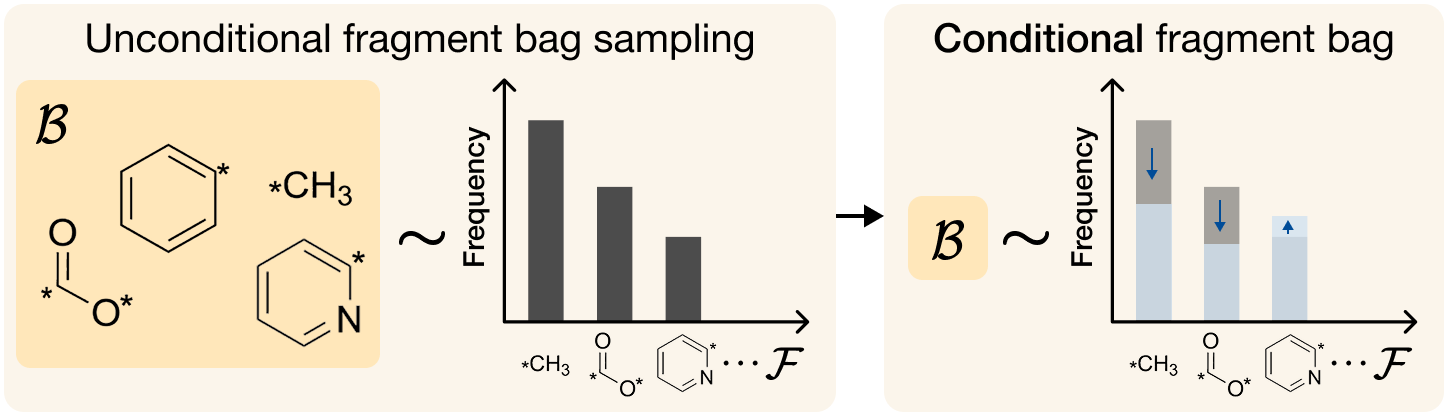
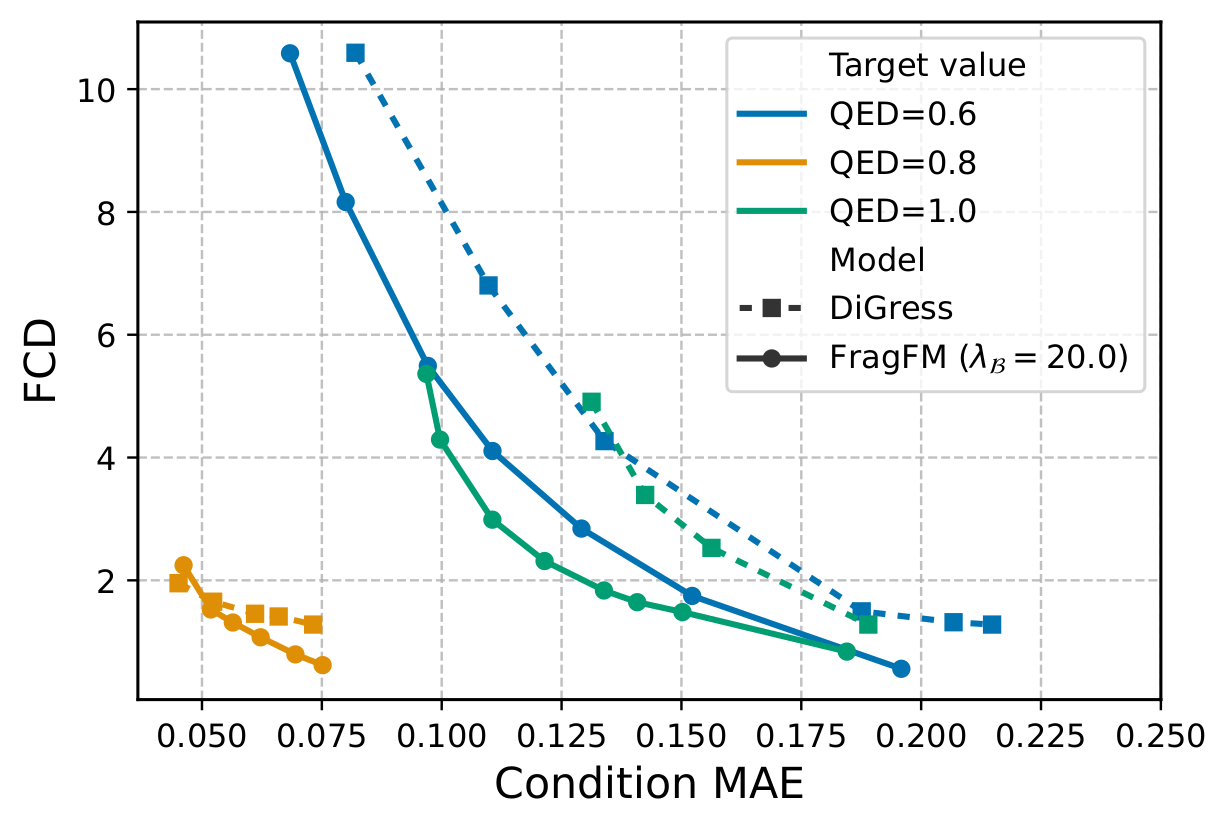
Both knobs drop out of a single Bayes decomposition of the property-conditioned in-bag transition kernel:
On the hard JAK2 docking-score task (top 0.08% of ZINC250k), bag guidance alone reaches the target while DiGress collapses in validity.
Synthesizability via Fragment-Bag Swap
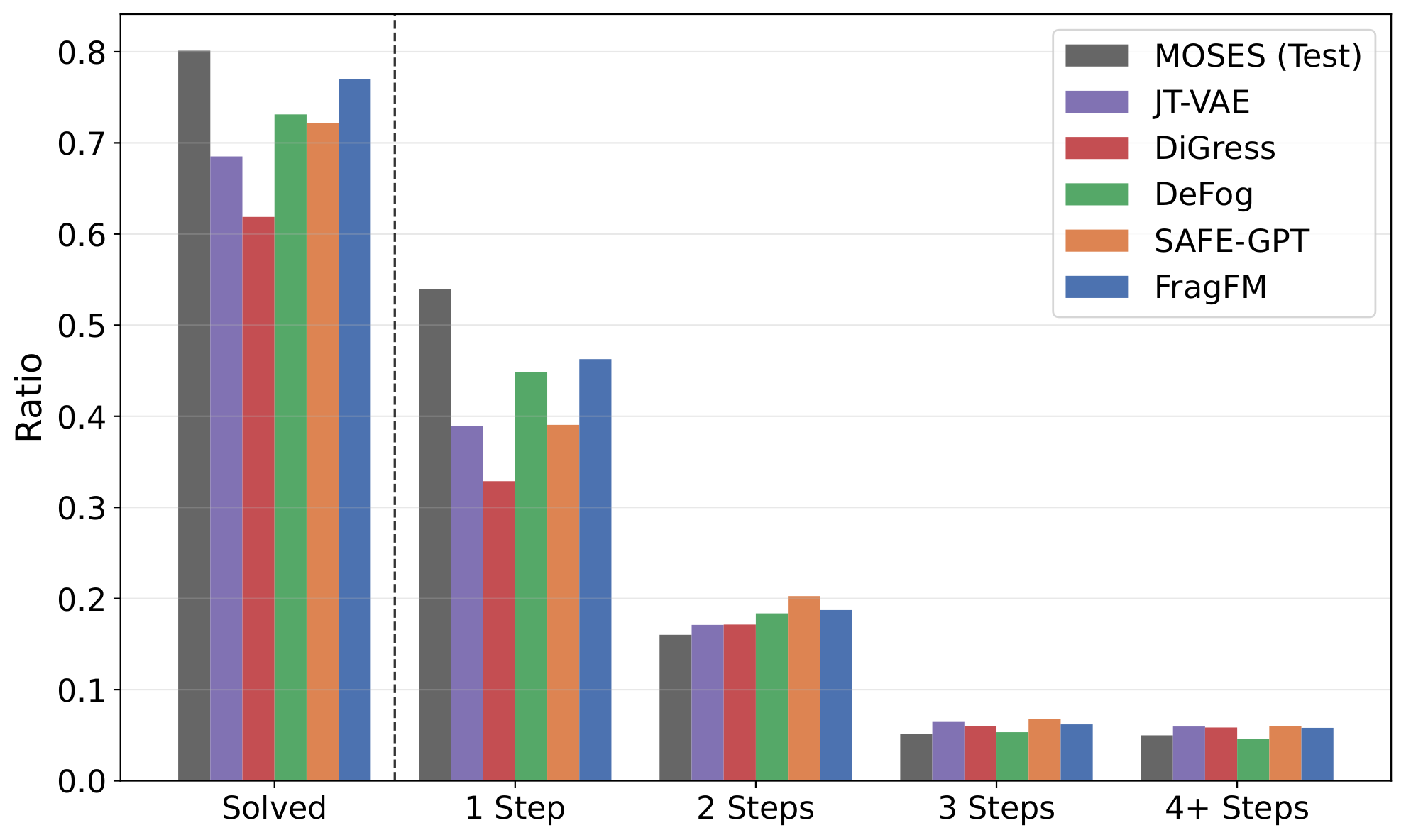
AIZynthFinder retrosynthesis on 25k MOSES samples: FragFM reaches 77% solved, close to the MOSES test set (80%). Swapping the fragment bag to fragments drawn from solved (synthesizable) molecules — no retraining — pushes it to 85%, exceeding the dataset itself.
| AIZynthFinder | Solved ↑ | 1 step | 2 steps | 3 steps | 4+ steps |
|---|---|---|---|---|---|
| MOSES (test set) | 80.1 | 53.9 | 16.0 | 5.17 | 5.0 |
| FragFM | 77.0 | 46.3 | 18.7 | 6.2 | 5.8 |
| FragFM · solved bag | 85.0 | 52.3 | 20.7 | 6.6 | 5.4 |
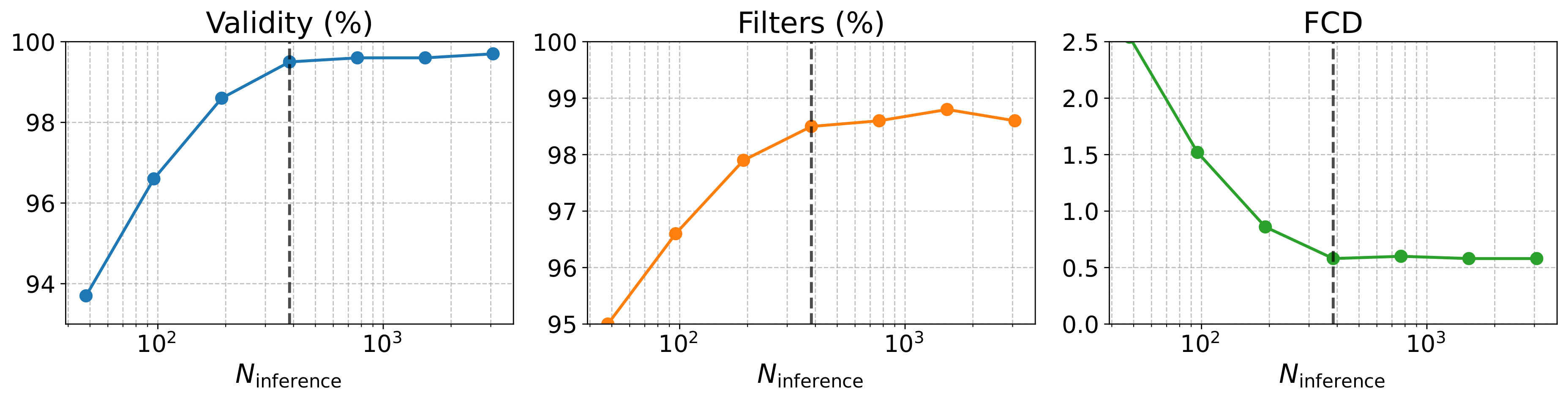
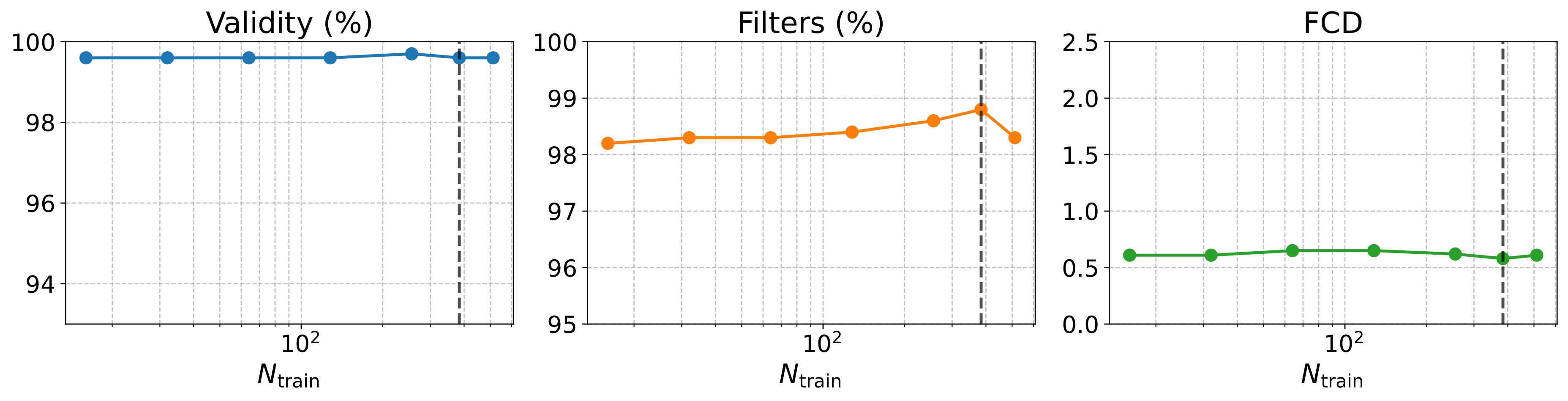
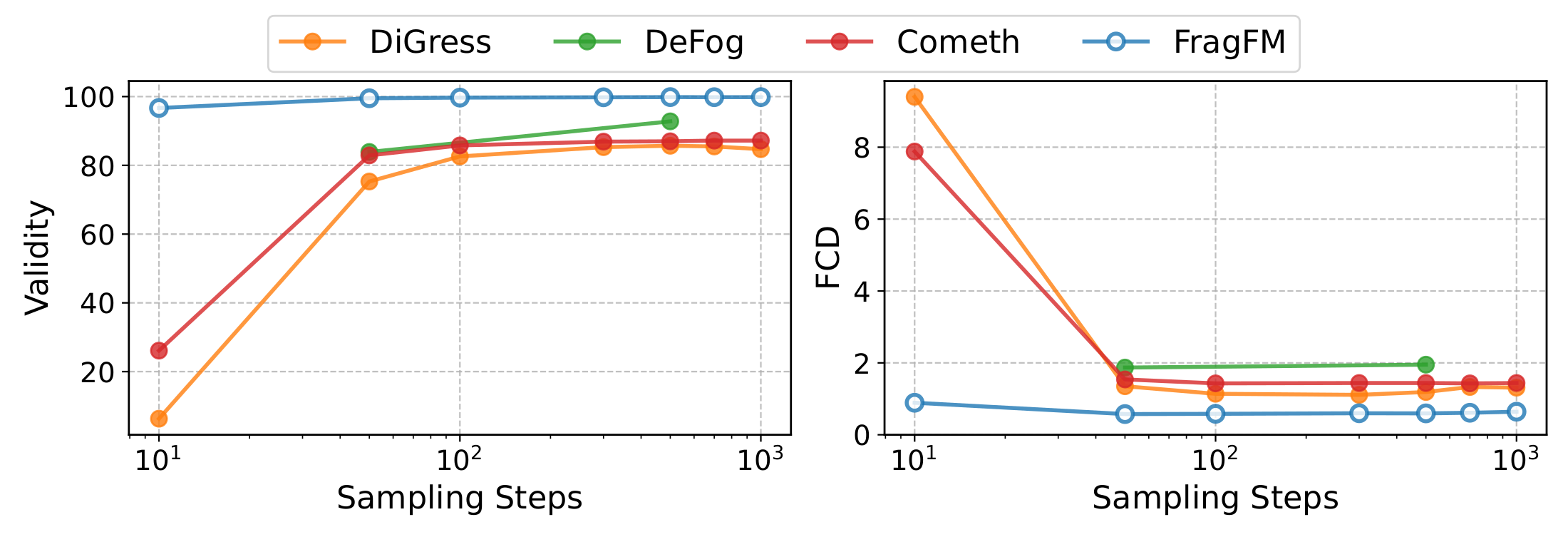
Scalability
Denoising at the fragment level — not over individual atoms — keeps validity above 95% and FCD below 1.0 even under aggressive step reduction, with ~5× faster wall-clock time than DiGress on NPGen.
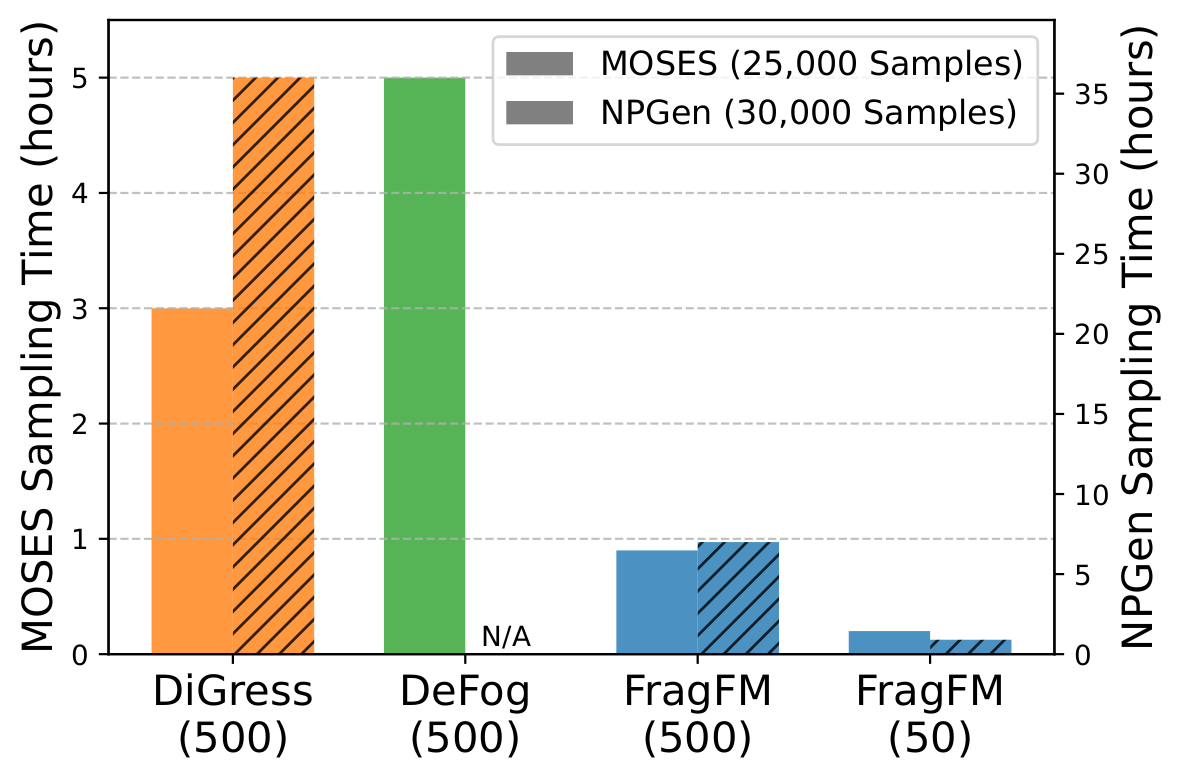
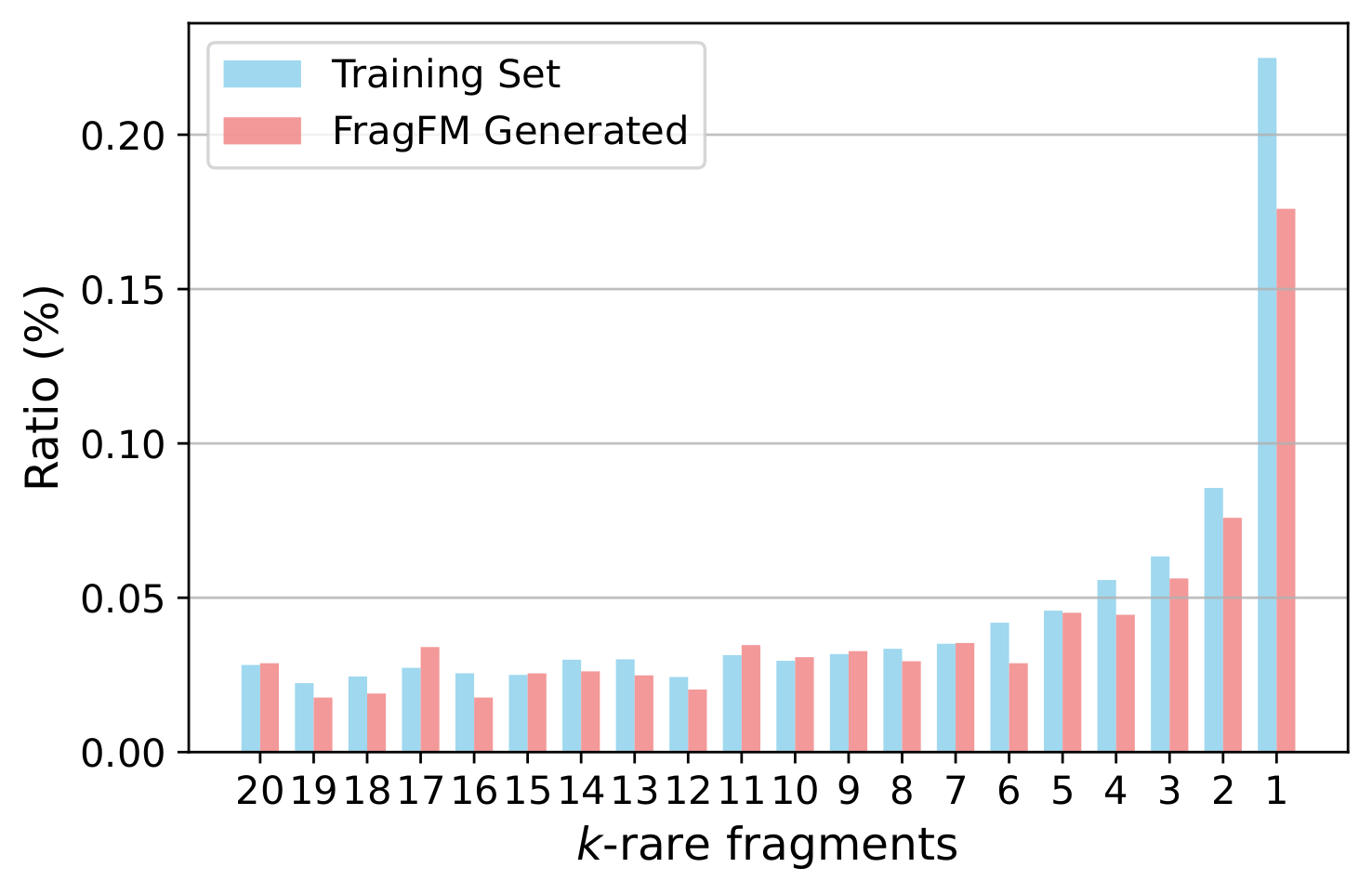
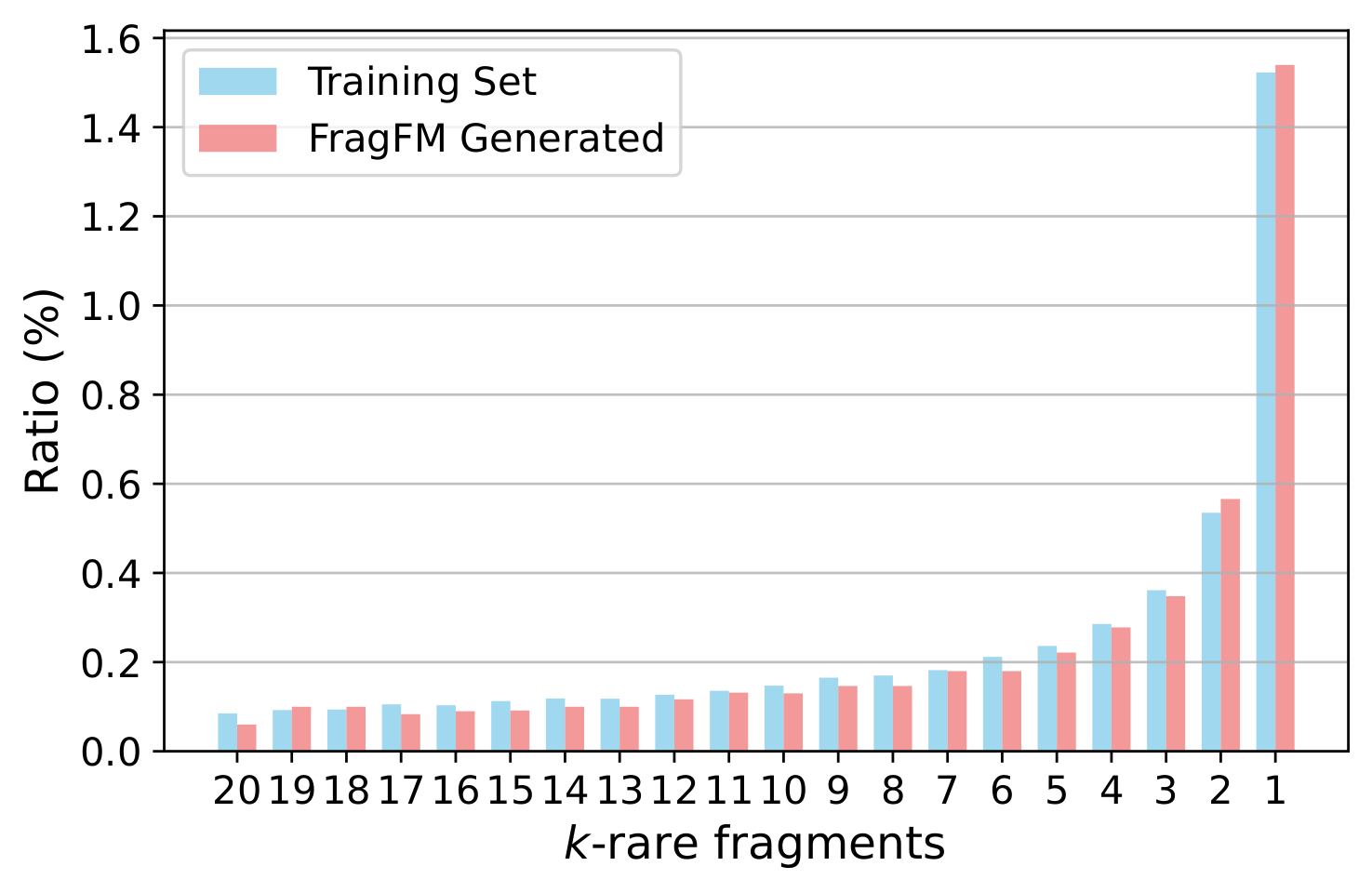
Additional Results
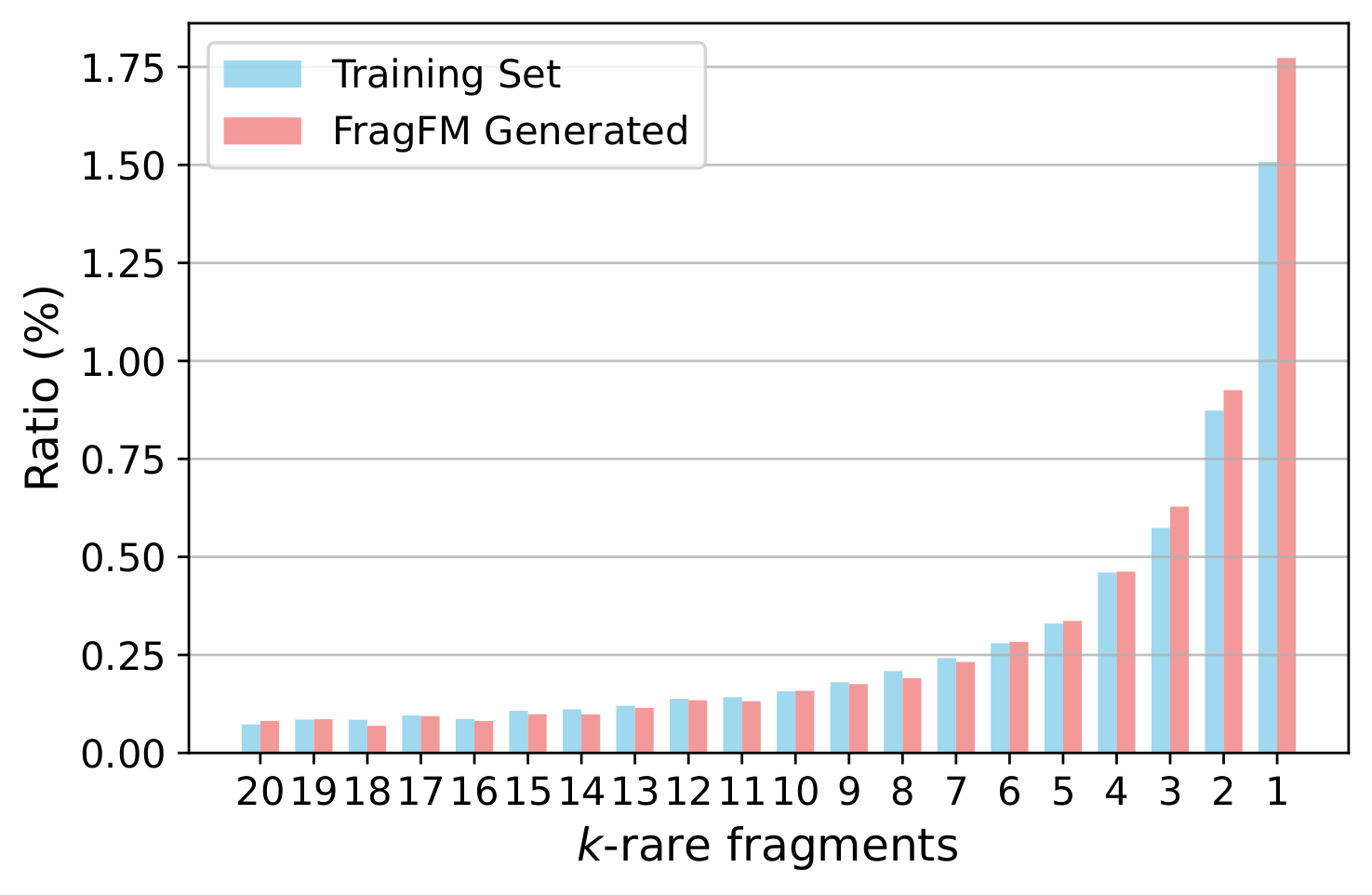
Ablations over the fragmentation rule, sampling temperature, fragment-bag size, and how well FragFM recovers rare fragments.
| Rule | MOSES | ZINC250k | |||||||
|---|---|---|---|---|---|---|---|---|---|
| Valid ↑ | Unique ↑ | Novel ↑ | Filters ↑ | FCD ↓ | SNN ↑ | Valid ↑ | NSPDK ↓ | FCD ↓ | |
| Training set | 100.0 | 100.0 | – | 100.0 | 0.48 | 0.59 | 100.0 | 0.0001 | 0.062 |
| BRICS (default) | 99.8 | 100.0 | 87.1 | 99.1 | 0.58 | 0.56 | 99.81 | 0.0002 | 0.630 |
| RECAP | 99.8 | 99.9 | 83.6 | 99.3 | 0.56 | 0.57 | 99.66 | 0.0003 | 0.580 |
| rBRICS | 99.8 | 100.0 | 88.5 | 98.7 | 0.58 | 0.56 | 99.79 | 0.0003 | 0.563 |
| 𝒯pred | 𝒯bag | Valid ↑ | Unique ↑ | Novel ↑ | Filters ↑ | FCD ↓ | SNN ↑ | Scaf ↑ |
|---|---|---|---|---|---|---|---|---|
| 1.0 | 1.0 | 99.8 | 100.0 | 87.1 | 99.1 | 0.58 | 0.56 | 10.9 |
| 1.0 | 1.5 | 99.2 | 100.0 | 94.2 | 98.3 | 0.90 | 0.52 | 13.5 |
| 1.5 | 1.0 | 99.7 | 100.0 | 88.6 | 98.8 | 0.88 | 0.54 | 11.0 |
| 1.5 | 1.5 | 99.2 | 100.0 | 94.5 | 98.3 | 0.91 | 0.51 | 13.1 |